 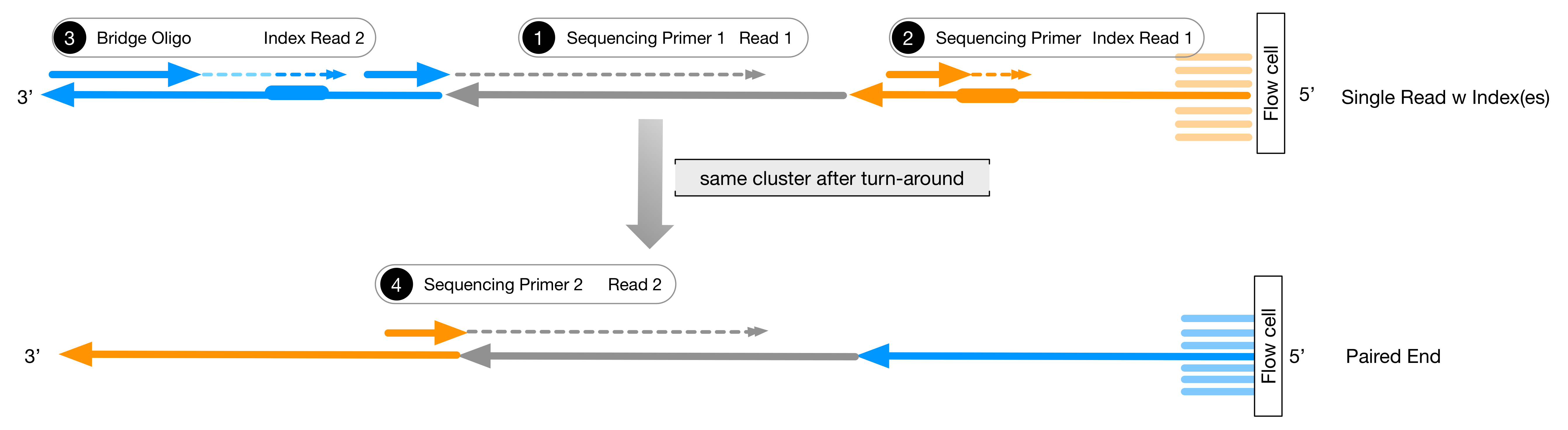
ArcherTM Illumina assays utilize dual-index barcoding to distinguish between samples. A fast and easy to use program to demultiplex amplicon pools with internal indexes is available at. generation sequencing on the MiSeq or HiSeq. A simple web page to design fusion primers compatible with iTru primers is available at. Thus, we show that these methods facilitate amplicon library construction for Illumina instruments at reduced cost with increased flexibility. We demonstrate the utility and versatility of our approaches with results from six projects using different implementations of our protocols. The number of sequencing reads from the amplicon pools can be adjusted, facilitating deep sequencing when required or reducing sequencing costs per sample to an economically trivial amount when deep coverage is not needed. Compatible with all KAPA Hyper library preparation kits, the HyperCap Workflow with Roche Universal Blocking Oligos, and PCR-free workflows. Sequences for AmpliSeq for Illumina Panels These CD and UD index adapters are arranged in the plate to enforce the recommended pairing strategy. The resulting quadruple-indexed amplicons have diversity at most base positions and can be pooled with any standard Illumina library for sequencing. High-quality adapters for multiplexed sequencing on 2- and 4-channel Illumina ® instruments, with patterned or non-patterned flow cells. The initial PCR products then serve as template for a second round of PCR with dual-indexed iTru or iNext primers (also used combinatorially) to make full-length libraries. These fusion primers can be used combinatorially to amplify samples within a 96-well plate (eight forward primers + 12 reverse primers yield 8 x 12 = 96 combinations), and the resulting amplicons can be pooled. 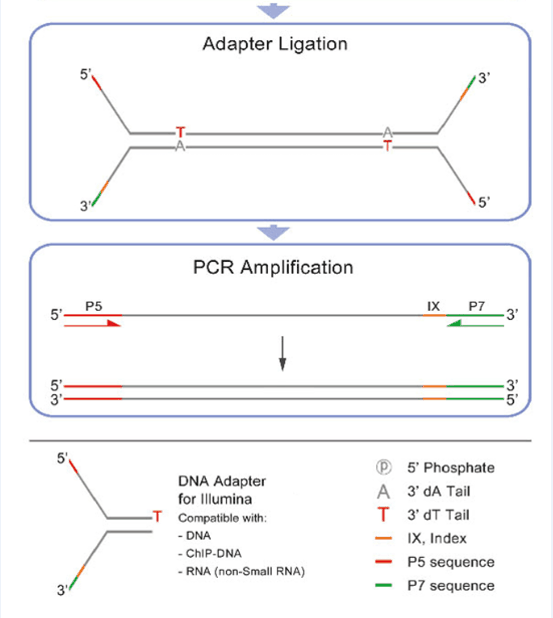
In the variant we use most frequently for large-scale projects, we fuse partial adapter sequences (TruSeq or Nextera) onto the 5’ end of locus-specific PCR primers with variable-length tag sequences between the adapter and locus-specific sequences. Here, we describe multiple ways to use the Adapterama system and other approaches for amplicon sequencing on Illumina instruments. Recommended tools would be for example these tools in their dedicated paired-end modes: BBduk, Skewer, HTStream, FASTP. This mode will not require any knowledge of the adapter sequences. In Adapterama I, we presented universal stubs and primers to produce thousands of unique index combinations and a modifiable system for incorporating them into Illumina libraries. Paired-end-read sequencing data should be trimmed using algorithms that make use of the paired-end nature to enable the most precise trimming. In many cases, NGS amplicon sequencing remains overly expensive and inflexible, with library preparation strategies relying upon the fusion of locus-specific primers to full-length adapter sequences with a single identifying sequence or ligating adapters onto PCR products. Next-generation sequencing (NGS) of amplicons is used in a wide variety of contexts. Illumina FASTQ file generation pipelines include an adapter trimming option for the removal of adapter sequences from the 3’ ends of reads. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |

 RSS Feed
RSS Feed
